 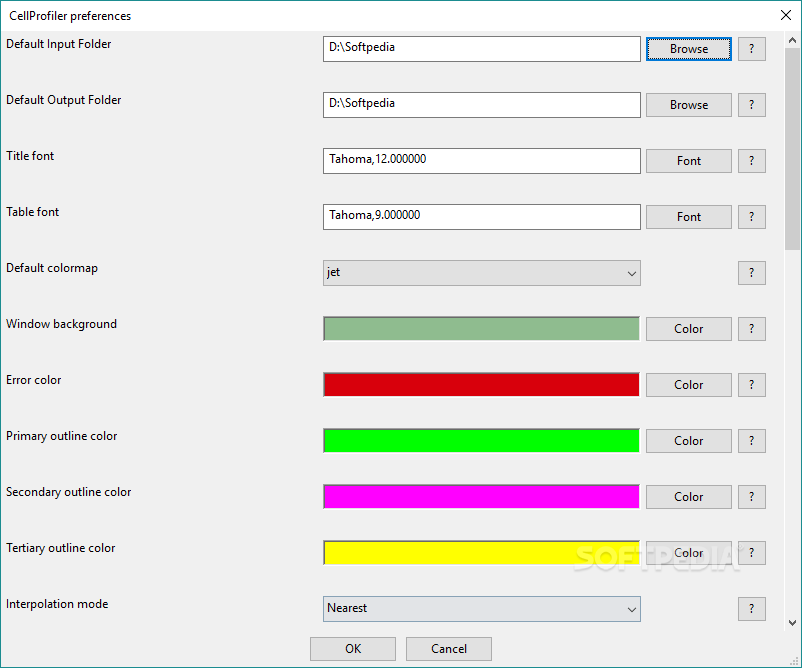
This pipeline shows how to input a color tissue image, split it into its component channels, and then identify individual cells from a particular stain and record the number of neighbors that each cell has. Here, we describe CellProfiler Analyst, open-source software for the interactive exploration and analysis of multidimensional data, particularly data from high-throughput, image-based. 
Every case can be easily to recreate the results. Download (0.4 MB) Tissue Neighbors: Tissue samples often have irregularly shaped cells with adjacent edges. The pipelines listed in the document along with the actual imagery are available as a part of plugin version. It contains the ready to use segmentation solution for a wide range of various imagery which includes:ĬellProfiler Analyst usage for advanced filtering In order to show that and also provide a quick starting point for users the Official user guide was prepared. Wide range of example usagesĭuring the testing phase of our plugin it turned out that combining parameter fitting and CellProfiler pipeline flow can result in a very flexible solution. See and run examples/use_cellstar.py as well as tests for more details. 
set_frame ( input_image ) segmentation, snakes = segmentator. Included is a supervised machine learning system which can be trained to recognize complex and subtle phenotypes, for automatic scoring of millions of cells. Segmentation ( segmentation_precision = 9, avg_cell_diameter = 35 ) segmentator. CellProfiler Analyst allows interactive exploration and analysis of data, particularly from high-throughput, image-based experiments. imread ( "input_images/sample_brightfield.tif" ) segmentator = cellstar. How to use package import cellstar input_image = imageio. CellProfiler Analyst allows the exploration and visualization of image-based data, together with the classification of complex. The plugin package includes not only the plugin itself but also examples of its usage to guide users on how to achieve best segmentation on a given type of imagery. Please visit our website for more details. The pipelines listed in the document along with the actual imagery are available as a part of plugin. This free software was originally designed by Broad Institute Imaging Platform. CellProfiler Analyst usage for advanced filtering. The important part of that solution is parameter fitting mechanism which allows to train and use CellStar for many different types of imagery. You can download CellProfiler-Analyst 2.2.1 from our software library for free. images from microfluidic chambers), however the algorithm can be also used to track other round objects (in brightfield as well as fluorescent images). It is optimized to segment and track images of budding yeast cells growing in monolayer (e.g. 
Since the segmentation and tracking of cells in brightfield images is considered to be a difficult and complex task, a number of software solutions have been already developed.ĬellStar is one of such algorithms. Free ImageJ image analysis software download, plugins, and documentation can all be found here. To limit manipulations during cell line preparation and phototoxicity during imaging, brightfield imaging is often considered. We’re open for innovation®, so visit us at KNIME.Automatic tracking of cells in time-lapse microscopy is required to investigate a multitude of biological questions. KNIME’s headquarters are based in Zurich, with additional offices in Konstanz, Berlin, and Austin. Orbit has been developed at Actelion Pharmaceuticals Ltd, now Idorsia Pharmaceuticals Ltd Carpenter and Thouis (Ray) Jones in the Sabatini Laboratory (Whitehead Institute for Biomedical Research) and Golland laboratory (MIT’s CSAIL) and is actively improved and maintained. The project was started in 2003 by Anne E. The CellProfiler project team is based in the Carpenter Lab at the Broad Institute of Harvard and MIT. The software was originally created at the Centre for Cancer Research & Cell Biology at Queen’s University Belfast, as part of research projects funded by Invest Northern Ireland and Cancer Research UK. QuPath is developed at the University of Edinburgh. Broad Institute Home Publications CellProfiler Analyst 3.0: Accessible data exploration and machine learning for image analysis. Eliceiri/LOCI group at the University of Wisconsin-Madison, and the Jug and Tomancak labs at the MPI-CBG in Dresden. CellProfiler Analyst 3.0: Accessible data exploration and machine learning for image analysis. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
